SERVICE
Single Cell RNA-Seq
Single Cell RNA-Seq
To overcome the limitations of transcriptome analysis of tissues or bulk cells, the diversity of each cellular gene expression can be identified by providing a single cell RNA-Seq service that analyze the transcripts of a single cell.
Service Feature
| Sample requirement |
Cell line, Primary cell, FACS sorted cell : 250,000 cells (Viability > 70%) Tissue : Over 200 mg of Fresh Tissue (in MACS buffer) |
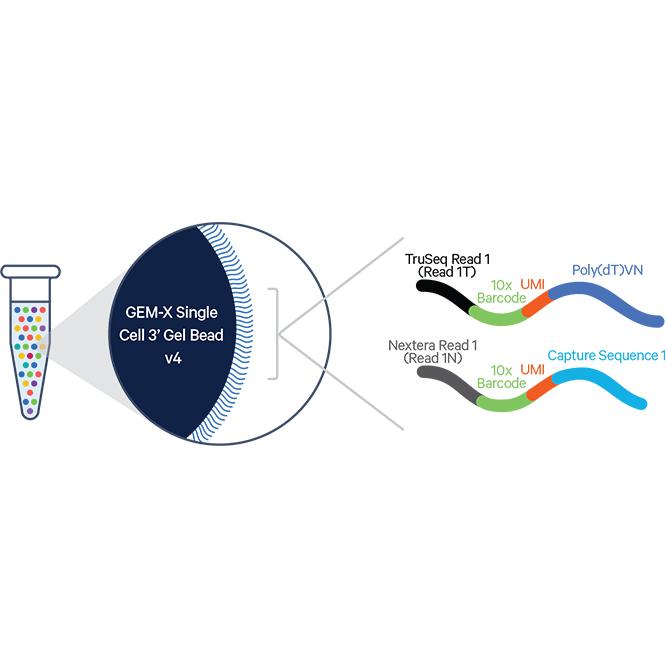
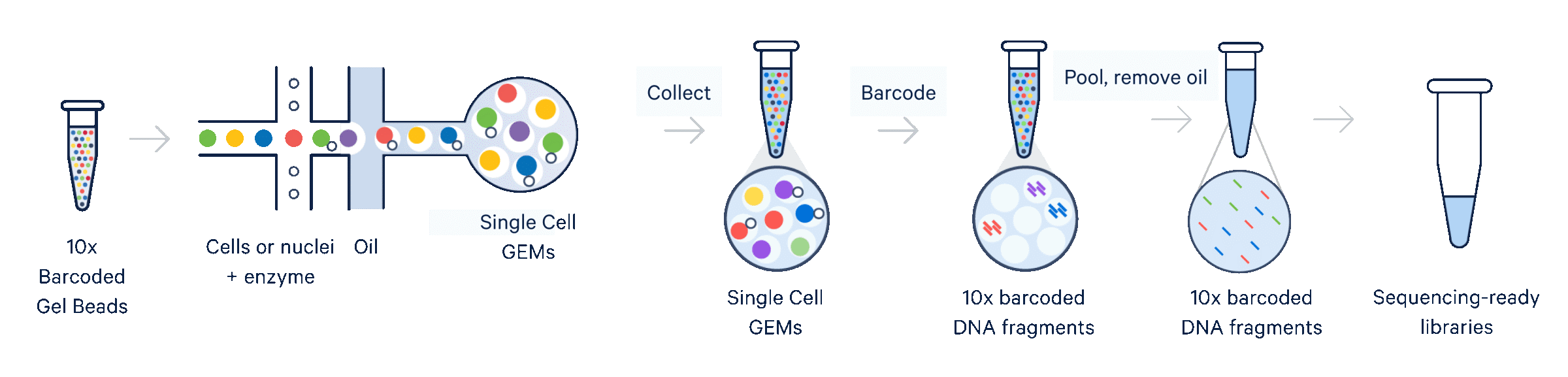
| Library method | 10X Genomics Next GEM-X 3’ technology |
| Number of Target cell | 3,000 ~ 20,000 cells |
| Turnaround Time | ~4 weeks after Cell counting & QC |
| Sample type | Cell, Tissue⁺ |
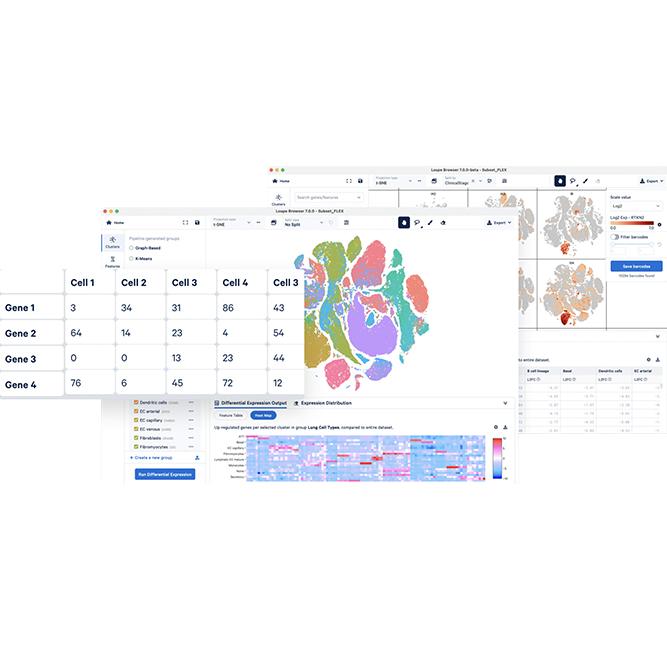
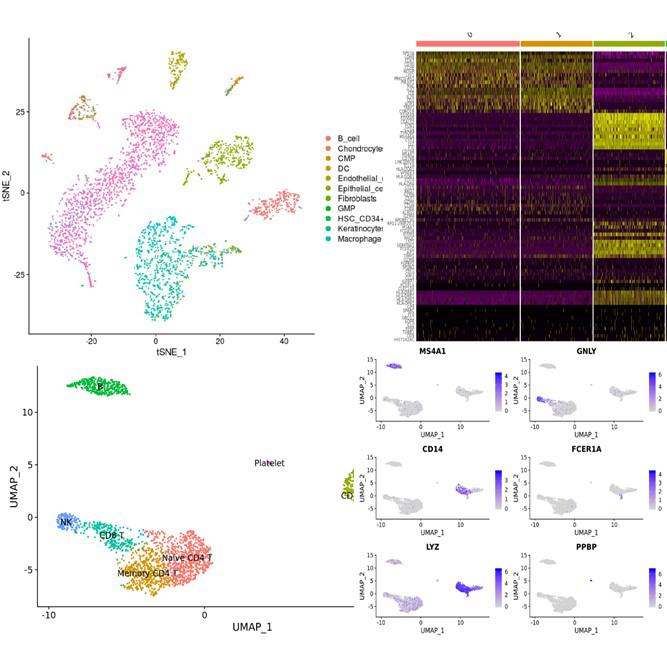
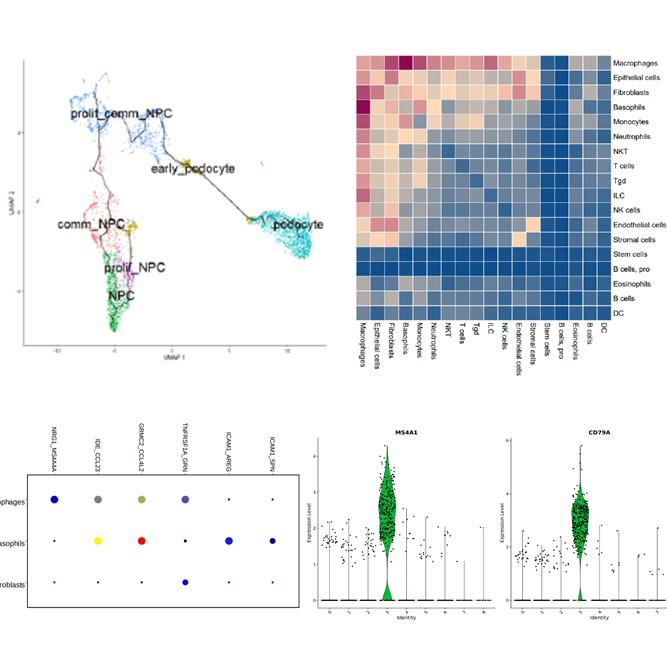
| Data analysis | ExSCEA Report, UMAP, t-SNE, PCA, Clustering heatmap (Seurat/Loupe based analysis) |
+ Tissue Dissociation process may require separate consultation.
Progress
-
1
Tissue dissociation
-
2
Cell QC.
-
3
GEM & Library prep.
-
4
NGS
-
5
Data Analysis & Report
Data Analysis
We provide 10X Genomics's basic support for Loupe Browser analysis of Cell Ranger output, and a variety of cluster-based comparative analysis and Cell type DE analysis through Excel based Single Cell Expression Analysis (ExSCEA) and WinSeurat (Windows based Seurat) developed and produced by Ebiogen.