SERVICE
DNA-Seq
Metagenome(16S V3-V4 Amplicon Seq.)
Provides metagenomic and microbiome analysis which identifies and quantifies microbial community (bacteria, fungi, etc) derived from various environments including soil, air, water, and human body by using 16s rRNA sequencing. V3-V4 Amplicon Sequencing is a service that analyzes the composition of microorganisms (Bacteria, Fungi, etc.) by sequencing the V3-V4 region of 16S rRNA. Since it only targets specific regions, it is suitable for analyzing samples in large quantities or for grasping the approximate composition of specific microbial communities. In this service, Bacteria sequenced the V3-V4 region and Fungi the ITS region to provide Metagenome and Microbiome analyses.
Service Feature
|
Sample requirement |
>0.1 ng/µl gDNA |
|
Library method |
16S rRNA Amplicon Library(16S/ITS) for Illumina |
|
NGS run format |
MiSeq, PE300bp |
|
Data yield |
~50,000 reads/sample |
|
Turnaround time |
~4 weeks after Library QC |
|
Sample type |
gDNA⁺ |
+ If you need gDNA preparation, you can proceed through consultation.
Progress
-
1
DNA QC
-
2
Library prep.
-
3
NGS
-
4
Data Analysis
-
5
Report
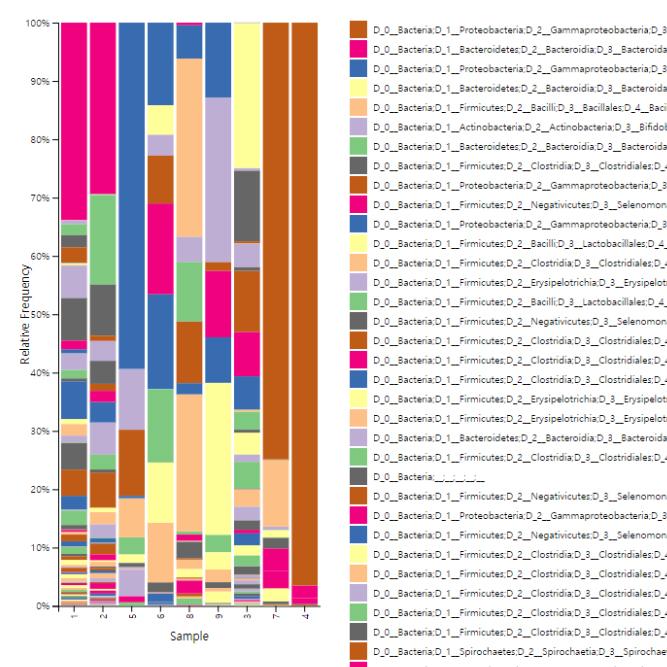
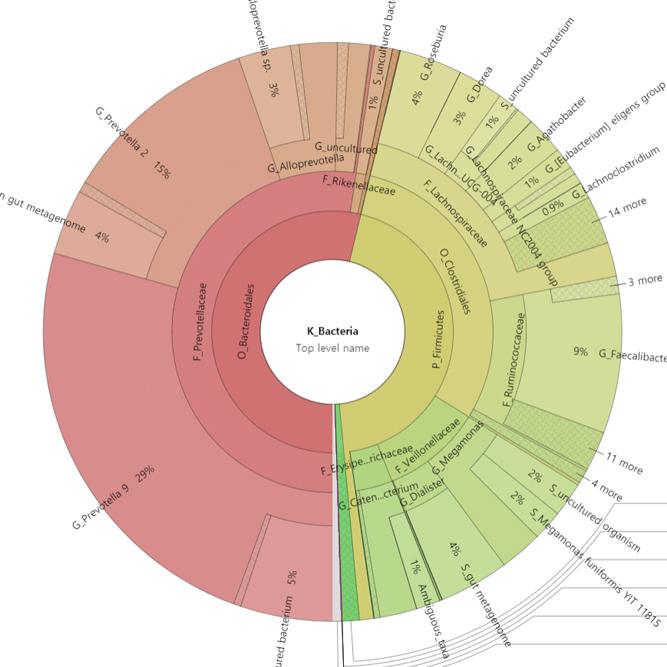
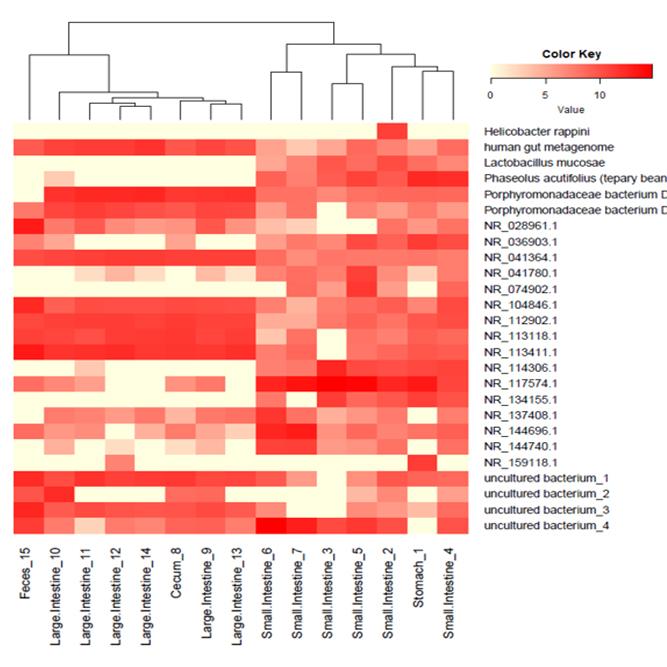
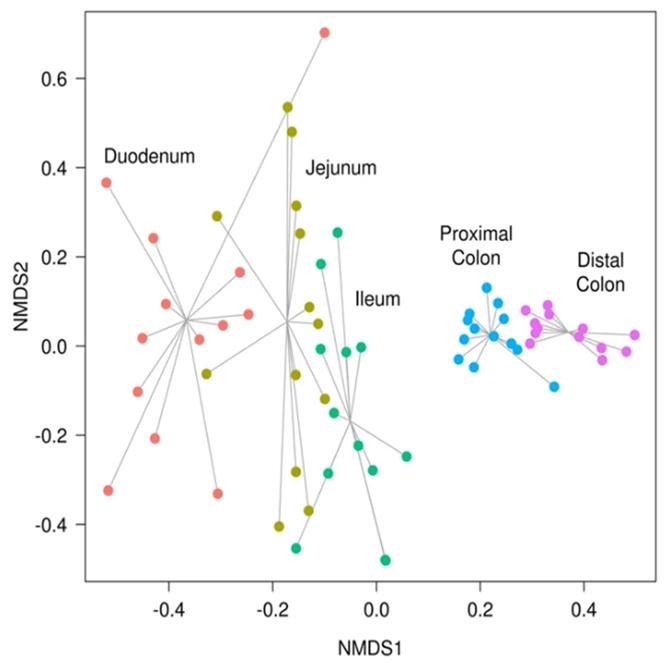
Data Analysis
We support ExMEGA report, various analysis such as Assembly Result, Feature Table, Taxonomy(Bar & Pie Chart, Profile Data), Diversity(Alpha, Beta), Sample Clustering et cetera.